Submissions






GlyTouCan
Glycan Structure Repository
GlyComb
Glycoconjugate Repository
GlycoPOST
Glycomics MS raw data RepositoryUniCarb-DR
Glycomics MS Repository for glycan annotations from GlycoWorkbench
LM-GlycoRepo
Repository for lectin-assisted multimodality dataAll Resources
Genes / Proteins / Lipids Glycans / Glycoconjugates Glycomes Pathways / Interactions / Diseases / OrganismsTools
Guidelines
MIRAGE G00033MO
G00033MO
Summary
- GlyTouCan ID
-
G00033MO
- IUPAC Condensed
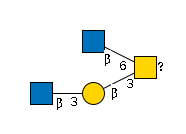
- Gal(b1-3)[GlcNAc(b1-6)]GalNAc(a1-
- Motifs
- O-Glycan core 2
- Subsumption Level
- Fully-defined saccharide
- Monoisotopic Mass
- 586.22
3D Structures
Organisms
| Organisms | Evidence |
|---|---|
| Mus musculus (house mouse) | |
| Rattus norvegicus (Norway rat) | |
| Homo sapiens (human) | |
| Severe acute respiratory syndrome coronavirus 2 | |
| Drosophila melanogaster (fruit fly) |
Taxonomic Hierarchy
GlycoGene Database (GGDB)
| Gene Symbol | Donor | Acceptor | Reducing terminal(Acceptor) | Product | Reducing terminal(Product) | Reference |
|---|---|---|---|---|---|---|
| B3GALNT2 | UDP-GalNAc |
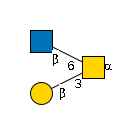 G00033MO
G00033MO
|
Ser/Thr |
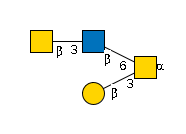 G01707KB
G01707KB
|
Ser/Thr | |
| GCNT1 | UDP-GlcNAc |
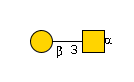 G00031MO
G00031MO
|
Ser/Thr (core 2) |
 G00033MO
G00033MO
|
Ser/Thr (core 2) | |
| GCNT3 | UDP-GlcNAc |
 G00031MO
G00031MO
|
Ser/Thr |
 G00033MO
G00033MO
|
Ser/Thr | |
| GCNT4 | UDP-GlcNAc |
 G00031MO
G00031MO
|
Ser/Thr |
 G00033MO
G00033MO
|
Ser/Thr |
| Gene Symbol | Donor | Acceptor | Reducing terminal(Acceptor) | Product | Reducing terminal(Product) | Reference |
|---|---|---|---|---|---|---|
| ST3GAL1 | CMP-Neu5Ac |
 G00033MO
G00033MO
|
O-glycan Synthesis |
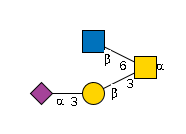 G85608AG
G85608AG
|
O-glycan Synthesis | |
| FUT2 | GDP-Fuc |
 G00033MO
G00033MO
|
O-glycan Synthesis |
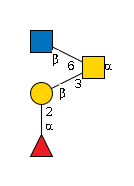 G32426JY
G32426JY
|
O-glycan Synthesis | |
| B4GALT4 | UDP-Gal |
 G00033MO
G00033MO
|
pNP |
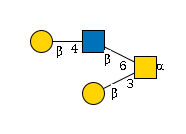 G64973KT
G64973KT
|
pNP | |
| B4GALT1 | UDP-Gal |
 G00033MO
G00033MO
|
O-glycan Synthesis |
 G64973KT
G64973KT
|
O-glycan Synthesis | |
| B3GALNT2 | UDP-GalNAc |
 G00033MO
G00033MO
|
(core2)-pNP |
 G01707KB
G01707KB
|
(core2)-pNP |
KEGG BRITE Database
Core Protein
| UniProt ID | Protein Name | Reference | Source |
|---|---|---|---|
| P01876 | Immunoglobulin heavy constant alpha 1 | ||
| P0DTC2 | Spike glycoprotein | ||
| P10761 | Zona pellucida sperm-binding protein 3 | ||
| P12259 | Coagulation factor V | ||
| P15941 | Mucin-1 | ||
| P20239 | Zona pellucida sperm-binding protein 2 | ||
| P21754 | Zona pellucida sperm-binding protein 3 | ||
| P35253 | Envelope glycoprotein | ||
| Q92954 | Proteoglycan 4 | ||
| Q9BYF1 | Angiotensin-converting enzyme 2 |
Sequence Descriptors
- GlycoCT
-
RES 1b:a-dgal-HEX-1:5 2s:n-acetyl 3b:b-dgal-HEX-1:5 4b:b-dglc-HEX-1:5 5s:n-acetyl LIN 1:1d(2+1)2n 2:1o(3+1)3d 3:1o(6+1)4d 4:4d(2+1)5n
- WURCS
- WURCS=2.0/3,3,2/[a2112h-1a_1-5_2*NCC/3=O][a2112h-1b_1-5][a2122h-1b_1-5_2*NCC/3=O]/1-2-3/a3-b1_a6-c1
Literature
| PubMed ID | Title | First Author | Publication Date | Source |
|---|---|---|---|---|
| 10753916 | Control of O-glycan branch formation. Molecular cloning and characterization of a novel thymus-associated core 2 beta1, 6-n-acetylglucosaminyltransferase | Schwientek T | 2000 Apr 14 |
|
| 9915862 | Molecular cloning and expression of a novel beta-1, 6-N-acetylglucosaminyltransferase that forms core 2, core 4, and I branches | Yeh JC | 1999 Jan 29 |
|
| 9988682 | Control of O-glycan branch formation. Molecular cloning of human cDNA encoding a novel beta1,6-N-acetylglucosaminyltransferase forming core 2 and core 4 | Schwientek T | 1999 Feb 19 |
|
| 10430883 | Expression cloning of a human alpha1, 4-N-acetylglucosaminyltransferase that forms GlcNAcalpha1-->4Galbeta-->R, a glycan specifically expressed in the gastric gland mucous cell-type mucin | Nakayama J | 1999 Aug 03 |
|
| 9530955 | Purification and characterization of the MUC1 mucin-type glycoprotein, epitectin, from human urine: structures of the major oligosaccharide alditols | Bhavanandan VP | 1998 Jan |
|
| 9857011 | Synthesis of poly-N-acetyllactosamine in core 2 branched O-glycans. The requirement of novel beta-1,4-galactosyltransferase IV and beta-1,3-n-acetylglucosaminyltransferase | Ujita M | 1998 Dec 25 |
|
| 1329093 | Expression cloning of a cDNA encoding UDP-GlcNAc:Gal beta 1-3-GalNAc-R (GlcNAc to GalNAc) beta 1-6GlcNAc transferase by gene transfer into CHO cells expressing polyoma large tumor antigen. | Bierhuizen MF | 1992 Oct 01 |
|
| 1627559 | The broad diversity of neutral and sialylated oligosaccharides derived from human salivary mucins | Klein A | 1992 Jul 7 |
|
| 1488058 | Quantitation and structures of oligosaccharide chains in human trachea mucin glycoproteins | Sangadala S | 1992 Dec 02 |
|
| 3345749 | Primary structure of neutral oligosaccharides derived from respiratory-mucus glycoproteins of a patient suffering from bronchiectasis, determined by combination of 500-MHz 1H-NMR spectroscopy and quantitative sugar analysis. 1. Structure of 16 oligosaccharides having the Gal beta(1----3)GalNAc-ol core (type 1) or the Gal beta(1----3)[GlcNAc beta(1----6)]GalNac-ol core (type 2) | Klein A | 1988 Feb 1 |
|
PubAnnotation
| Title | Authors | Source | Publication Date | PubMed ID |
|---|---|---|---|---|
| Recombinant MUC1 probe authentically reflects cell-specific O-glycosylation profiles of endogenous breast cancer mucin. High density and prevalent core 2-based glycosylation. |
|
J Biol Chem | 2002 Jul 19 | 12000758 |
| Specificities of N-acetylglucosamine-6-O-sulfotransferases in relation to L-selectin ligand synthesis and tumor-associated enzyme expression. |
|
J Biol Chem | 2002 Feb 8 | 11726653 |
| The relative activities of the C2GnT1 and ST3Gal-I glycosyltransferases determine O-glycan structure and expression of a tumor-associated epitope on MUC1. |
|
J Biol Chem | 2001 Apr 6 | 11118434 |
| MUC1 and cancer. |
|
Biochim Biophys Acta | 1999 Oct 8 | 10571020 |
| An alpha2,3 sialyltransferase (ST3Gal I) is elevated in primary breast carcinomas. |
|
Glycobiology | 1999 Dec | 10561455 |
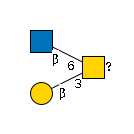 G00034MO
G00034MO
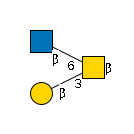 G24720OO
G24720OO
 G27236KE
G27236KE
